This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein interaction networks?
When studying a disease or gene, it is important to study protein-protein interactions (PPIs) because proteins function via interactions with other proteins, and a single protein's function can change depending on the other proteins involved. Results from many experiments to determine protein-protein interactions can be combined to create protein interaction networks, which provide a large web of interacting proteins. Online tools that combine primary paper's data to show protein interaction networks are a valuable way learn about a protein of interest. One of these online tools is called STRING, which combines data from high-throughput lab experiments, proteins that are conserved and co-expressed, and previous knoweldge from other databases (such as IntAct or BioGRID), along with genomic content predictions and automated textmining to create protein interaction networks [1]. Protein interaction networks are helpful when studying a specific protein because you can learn pathways the protein of interest is involved in or choose to test whether a predicted interaction actually occurs.
STRING, BioGRID, and IntAct are three online tools with information about protein interactions.
What proteins interact with AIRE?
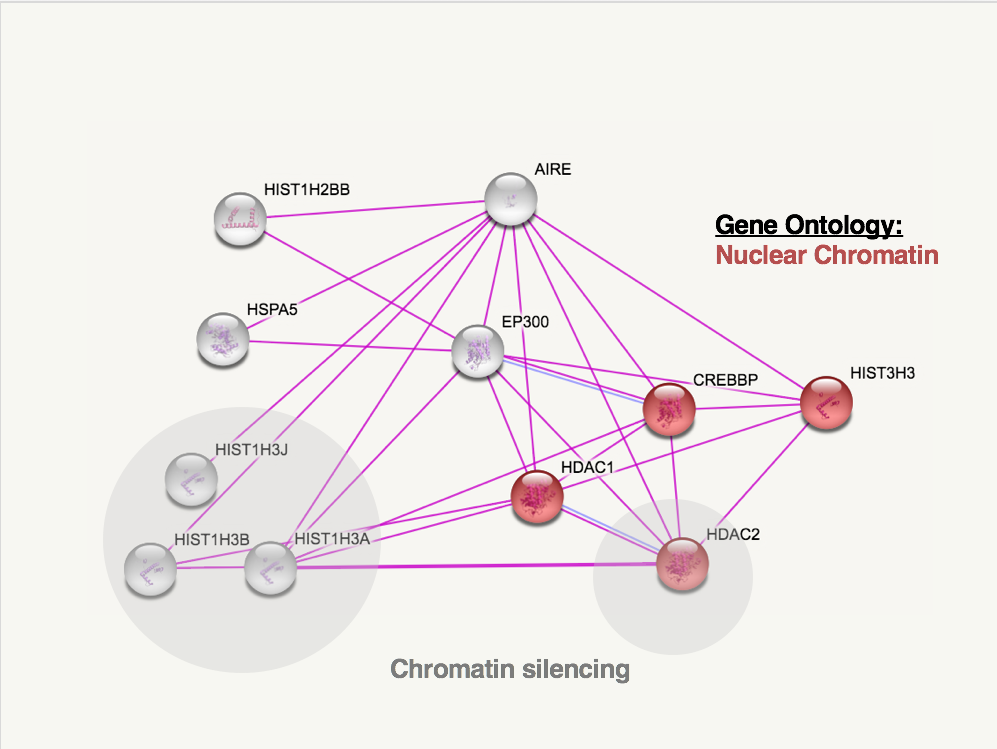
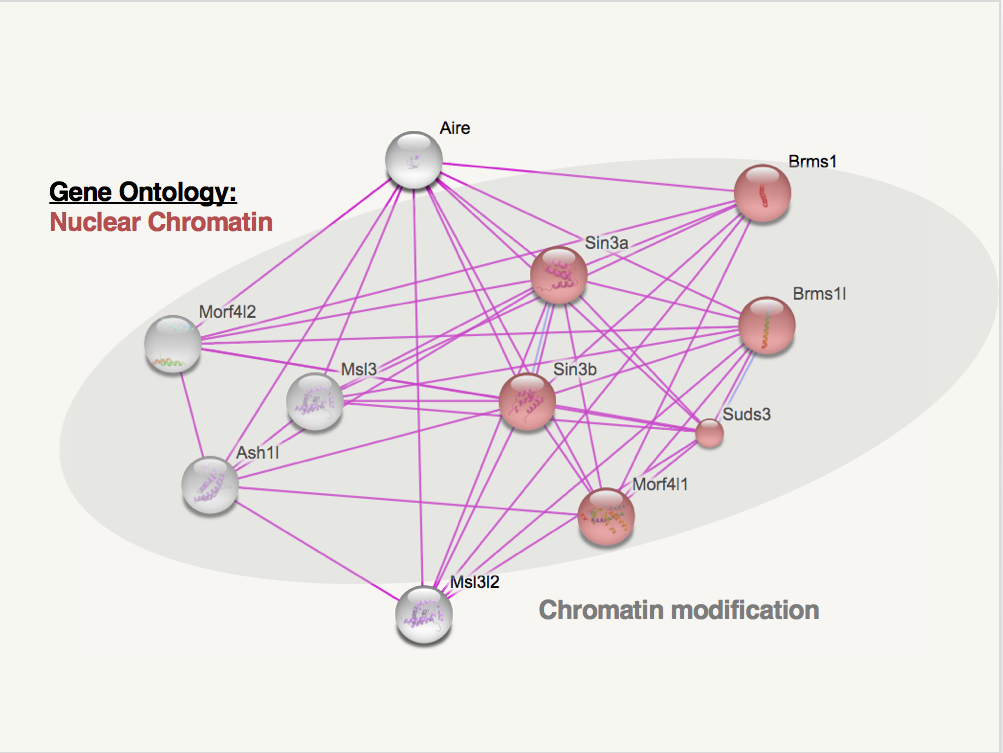
As a transcription factor, AIRE interacts with many different proteins, including DNA and chromatin binding proteins. Many of the proteins in AIRE's protein interaction network have biological process gene ontology annotation related to transcription, such as positive regulation of transcription or DNA-templated transcription initiation from RNA polymerase II promoter [2]. The images below show human and mouse AIRE protein interaction networks from STRING. These interaction networks only display proteins known to interact with AIRE from experimental data, and only 10 interacting proteins are shown in each map.
Analysis
Two gene ontology groups are noted above are important for AIRE's role as a transcription factor. The interacting proteins in red are categorized as "nuclear chromatin" for gene ontology's cellular component. It makes sense that AIRE interacts with so many proteins involved with nuclear chromatin because AIRE is a transcription factor that causes the expression of proteins that are usually tissue-specific. In other words, AIRE's activity in the thymus and extra thymic AIRE expressing cells leads to the expression of proteins that would be condensed in the thymic or extra thymic cells becuase they are normally only expressed in other, specific tissues.
The grey circles indicate proteins involved with chromatin silencing in humans and chromatin modification in mice. Again, like the nuclear chromatin ontology, it makes sense that AIRE interacts with proteins involved in chromatin because AIRE's role is to recruit a transcription complex to areas of the genome that are not normally expressed in thymic/extra thymic AIRE expressing cells. The proteins AIRE expresses are then presented to immune cells which are killed if they bind tightly to these self-antigens that AIRE expressed. This process is beneficial to ensure that immune cells do not attack bodily proteins.
The grey circles indicate proteins involved with chromatin silencing in humans and chromatin modification in mice. Again, like the nuclear chromatin ontology, it makes sense that AIRE interacts with proteins involved in chromatin because AIRE's role is to recruit a transcription complex to areas of the genome that are not normally expressed in thymic/extra thymic AIRE expressing cells. The proteins AIRE expresses are then presented to immune cells which are killed if they bind tightly to these self-antigens that AIRE expressed. This process is beneficial to ensure that immune cells do not attack bodily proteins.
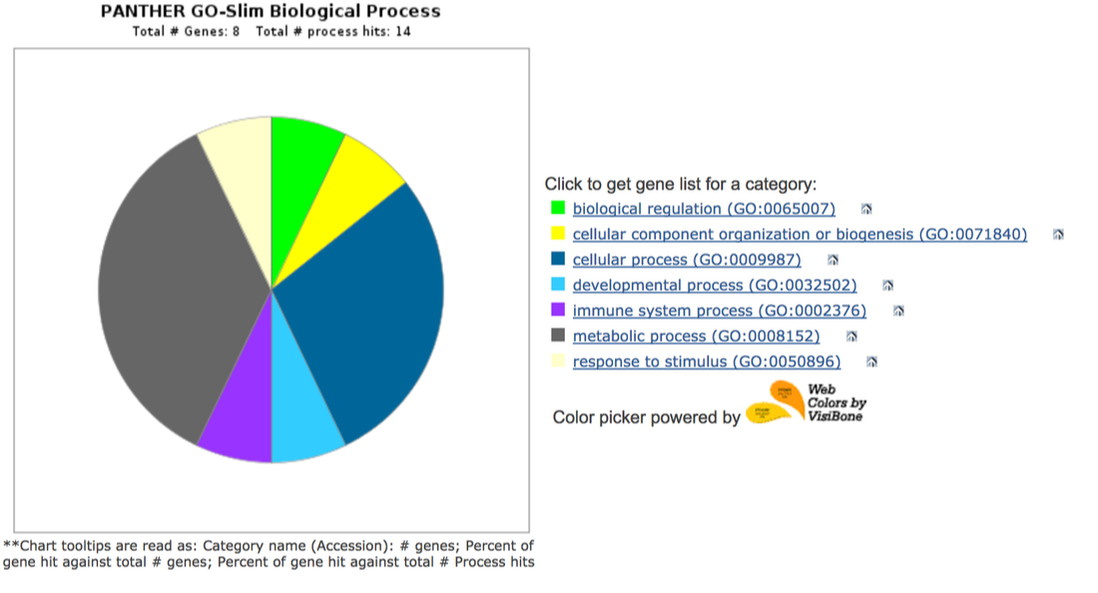
Panther Gene Ontology Data
In general, the gene ontology of proteins that interact with AIRE in mice and humans are very similar. The above image shows the biological processes of proteins that interact with AIRE. As shown in the pie chart, many are involved in metabolic processes, biological regulation, cellular processes, immune system processes, and developmental processes. The molecular functions of these proteins can be seen above in the interaction network, but many are involved in chromatin binding and transcription factor binding. These molecular and biological functions make sense given AIRE's role in transcriptional activation.
References:
[1] STRING Database--Content. http://string-db.org/cgi/about.pl?UserId=lse9kRZ202nG&sessionId=9FkzvVG7Hc9y&footer_active_subpage=content
[2] STRING Output AIRE Homo sapien. http://string-db.org/cgi/network.pl?taskId=tFh8CAumBGpw
[3] STRING Output AIRE Mus musculus. http://string-db.org/cgi/network.pl?taskId=BpvwmRK0XHbN
[1] STRING Database--Content. http://string-db.org/cgi/about.pl?UserId=lse9kRZ202nG&sessionId=9FkzvVG7Hc9y&footer_active_subpage=content
[2] STRING Output AIRE Homo sapien. http://string-db.org/cgi/network.pl?taskId=tFh8CAumBGpw
[3] STRING Output AIRE Mus musculus. http://string-db.org/cgi/network.pl?taskId=BpvwmRK0XHbN